Download Exercise 1
Transcript
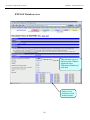
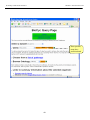
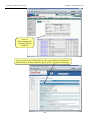
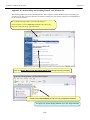
Workshop on Bacterial Genomics Module 1: Artemis Artemis Exercise 2 This exercise will look at a section of the Malaria genome. You will need to close down the last Artemis exercise if you haven’t already done so. Then start a new Artemis Session, as before, using the file ‘Malaria.embl’ in the current directory (Module_2_Artemis). Unlike the Salmonella exercise, in this instance the annotation and sequence are contained within the same file ‘Malaria.embl’ The sequence you are going to look at is a small region of contrived sequence (~21 kb) taken from Plasmodium falciparum chromosome 13. You will see 7 CDSs, some with multiple exons. As a gentle introduction to splicing we would like you to look at the genes named , PF13_0119, MAL13P1.294 and PF13_0061. They have only been partially characterised and may in fact be missing exons. Have a look at these CDSs and confirm, edit or dismiss the proposed gene models by using G+C content, database searches and looking for splice sites (Appendix IX). G+C content is a very good indicator of coding capacity in Malaria. On average, the coding regions are ~23% G+C and the non-coding regions are ~19%. Have a look at the G+C content for this region by selecting the appropriate graph. Left click within the graph window and then select by clicking on the exons to see how this relates to the G+C peaks on the graph. Note, we will cover the principals and methods of gene prediction in much more detail in a module 3. fasta banner -25-