Download CLC bio
Transcript
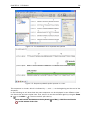
CHAPTER 2. TUTORIALS 45 Suppose you are working with the ATP8a1 protein sequence which is a phospholipid-transporting ATPase expressed in the adult house mouse, Mus musculus. To obtain more information about this molecule you wish to query the peptides held in the Swiss-Prot* database to find homologous proteins in humans Homo sapiens, using the Basic Local Alignment Search Tool (BLAST) algorithm. This tutorial involves running BLAST remotely using databases housed at the NCBI. Your computer must be connected to the internet to complete this tutorial. 2.5.1 Performing the BLAST search Start out by: select protein ATP8a1 | Toolbox | BLAST Search ( ) | NCBI BLAST ( ) In Step 1 you can choose which sequence to use as query sequence. Since you have already chosen the sequence it is displayed in the Selected Elements list. Click Next. In Step 2 (figure 2.12), choose the default BLAST program: blastp: Protein sequence and database and select the Swiss-Prot database in the Database drop down menu. Figure 2.12: Choosing BLAST program and database. Click Next. In the Limit by Entrez query in Step 3, choose Homo sapiens[ORGN] from the drop down menu to arrive at the search configuration seen in figure 2.13. Including this term limits the query to proteins of human origin. Choose to Open your results. Click Finish to accept the parameter settings and begin the BLAST search. The computer now contacts NCBI and places your query in the BLAST search queue. After a short while the result should be received and opened in a new view.