Download CLC Sequence Viewer
Transcript
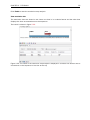
CHAPTER 9. VIEWING AND EDITING SEQUENCES 103 Figure 9.11: Creating a sequence. • Latin name. The Latin name for the species. • Type. Select between DNA, RNA and protein. • Circular. Specifies whether the sequence is circular. This will open the sequence in a circular view as default. (applies only to nucleotide sequences). • Description. A description of the sequence. • Keywords. A set of keywords separated by semicolons (;). • Comments. Your own comments to the sequence. • Sequence. Depending on the type chosen, this field accepts nucleotides or amino acids. Spaces and numbers can be entered, but they are ignored when the sequence is created. This allows you to paste (Ctrl + V on Windows and + V on Mac) in a sequence directly from a different source, even if the residue numbers are included. Characters that are not part of the IUPAC codes cannot be entered. At the top right corner of the field, the number of residues are counted. The counter does not count spaces or numbers. Clicking Finish opens the sequence. It can be saved by clicking Save ( of the sequence view into the Navigation Area. 9.7 ) or by dragging the tab Sequence Lists The Sequence List shows a number of sequences in a tabular format or it can show the sequences together in a normal sequence view. Having sequences in a sequence list can help organizing sequence data. The sequence list may originate from an NCBI search (chapter 10.1). Moreover, if a multiple sequence fasta file is imported, it is possible to store the data in a sequences list. A Sequence List can also be generated using a dialog, which is described here: select two or more sequences | right-click the elements | New | Sequence List ( This action opens a Sequence List dialog: )