Download PDF version
Transcript
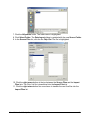
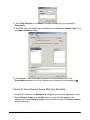
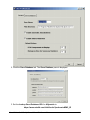
Normalization Relative to Positive Controls Across Experiments This is done by dividing the values in the dataset by the median value of the positive controls across all samples. For example: Gene i sample j /median (all positive control genes across all samples) Gene i sample k / median (all positive control genes across all samples) Refer to the above image. Actions 1. Click a complete dataset in the Experiments navigator. The item is highlighted. 2. Click the Normalize toolbar icon , or select Normalize from the Data menu, or rightclick the item and select Normalize from the shortcut menu. The first Normalization dialog is displayed. 3. Double-click the Positive and Negative Control Genes radio button, or click it and click Next. The second Normalization dialog is displayed. GeneLinker Gold 3.1 / GeneLinker Platinum 2.1 270
Related documents

Apple Power Macintosh 9600 Technical information
D ataRa ay Inc c.

Apple Macintosh Performa 6360 Technical information

ScanArray Express Microarray Analysis System User Manual

the user manual

DPoE™ 1GIG ™ Power Patch Panel

RFXpert Manual V1 1.2

Engene User Manual

TIGR MultipleExperimentViewer (MEV)

User`s Manual

TIBCO Spotfire Lead Discovery 2.1 – User`s Manual

USER MANUAL ENGLISH Version 2.11
Wood Working Plans mold 8 (1)

TIBCO Spotfire DecisionSite 9.1.1 Statistics

lire - Asti

SIEVE 1.1.0 User Guide Version B

321 Studios Meridian Internet Telephony Gateway (ITG) Trunk 2.0 User's Manual