Download PDF version
Transcript
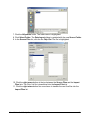
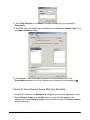
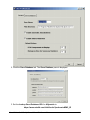
Tutorial 3A: Step 2 Perform Partitional Clustering Perform Partitional Clustering 1. If the new Norm: Scaled min to max dataset in the Experiments navigator is not already highlighted, click it. 2. Click the Partitional Clustering toolbar icon , or select Partitional Clustering from the Clustering menu, or right-click and select Partitional Clustering from the shortcut menu. The Partitional Clustering parameters dialog is displayed. 3. Set dialog parameters. Parameter Clustering Orientation Distance Measurements: Between Data Points Algorithm Properties: Type Algorithm Properties: Neighbors to Examine Algorithm Properties: Neighbors in Common Setting Cluster Genes Euclidean Jarvis-Patrick 6 2 4. Click OK. The clustering operation is performed and upon successful completion, a new J-P (6,2): genes | Euclid | average experiment is added to the Experiments navigator under the original dataset. If you have automatic visualizations enabled in your user preferences, a matrix tree plot of the clustering results is displayed. Tutorial 3A: Step 3 Create a Matrix Tree Plot If the matrix tree plot is already displayed, there is no need to recreate it. Read the sections below the image for information about the plot. GeneLinker Gold 3.1 / GeneLinker Platinum 2.1 80
Related documents

Apple Power Macintosh 9600 Technical information
D ataRa ay Inc c.

Apple Macintosh Performa 6360 Technical information

ScanArray Express Microarray Analysis System User Manual

the user manual

DPoE™ 1GIG ™ Power Patch Panel

RFXpert Manual V1 1.2

Engene User Manual

TIGR MultipleExperimentViewer (MEV)

User`s Manual

TIBCO Spotfire Lead Discovery 2.1 – User`s Manual

USER MANUAL ENGLISH Version 2.11
Wood Working Plans mold 8 (1)

TIBCO Spotfire DecisionSite 9.1.1 Statistics

lire - Asti

SIEVE 1.1.0 User Guide Version B

321 Studios Meridian Internet Telephony Gateway (ITG) Trunk 2.0 User's Manual